
ChemComm 2020
Non-canonical lipoproteins with programmable assembly and architecture

Langmuir 2020
Ion transport through the tight junction pores formed by transmembrane claudin proteins between membranes of two adjacent cells. The ion conductance is regulated by the electrostatic environment created by the pore-lining amino acid residues
Physical Chemistry Chemical Physics 2020
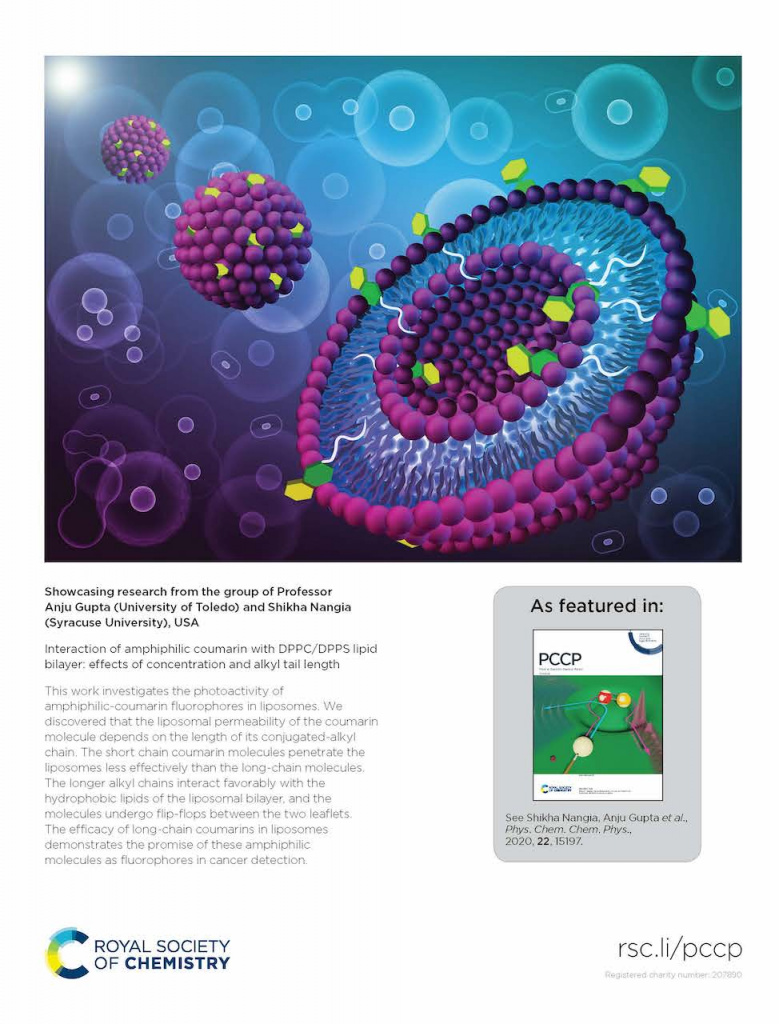
A Tale of Two Tails: Photoactivity through amphiphilicity. We discovered that the liposomal permeability of the coumarin molecule depends on the length of its conjugated-alkyl chain. The short-chain coumarin molecules penetrate the liposomes less effectively than the long-chain molecules. The longer alkyl chains interact favorably with the hydrophobic lipids of the liposomal bilayer, and the molecule undergo flip-flops between the two leaflets. The efficacy of long-chain coumarins in liposomes demonstrates the promise of these amphiphilic molecules as fluorophores in cancer detection.

Journal of Chemical Theory and Computation 2019
The protein association energy landscape (PANEL) method provides an extensive data set of membrane protein association in lipid milieu using a combination of stochastic sampling and equilibration simulations. The results are obtained in a fraction of the simulation cost compared to self-assembly simulations

The Journal of Physical Chemistry 2018
Molecular architecture of the claudin tight junction proteins. The claudin family of transmembrane proteins forms tight junctions between the adjacent cells in epithelial and endothelial tissues. These tight junctions control the transport and permeability of ions and small molecules through the paracellular space. The tight junction permeability is critical in understanding physiological processes such as the blood–brain barrier, renal filtration, ionic hemostasis, and drug transport across tissue barriers. This multiscale dynamics study of classic claudins provides the molecular architecture of the tight junction pores as well as the putative conformation of claudin proteins that acts as a receptor for the Clostridium perfringens enterotoxin.
Langmuir 2018
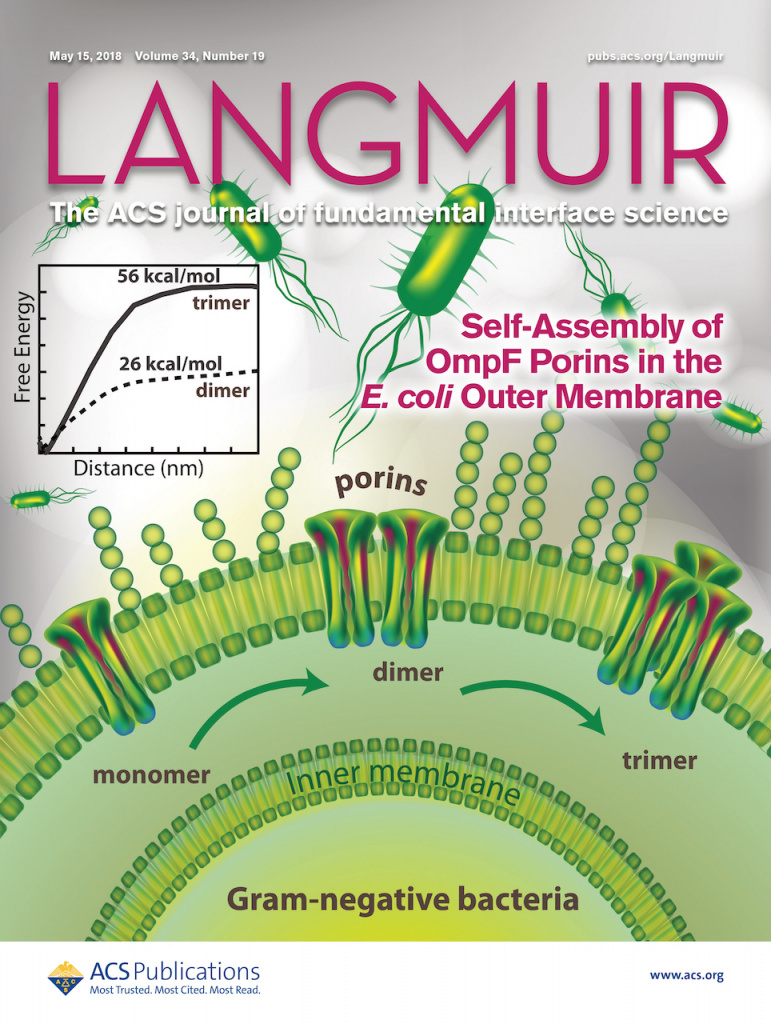
The channel-forming protein OmpF porin from Escherichia coli spans the bacterial outer membrane and facilitates the diffusion-mediated influx of nutrients into the cell. Using molecular dynamics approach, we explore the thermodynamics of OmpF assembly over microsecond timescale simulations, enabled by the newly developed coarse-grain force field parameter set for E. coli outer membrane lipids.

The Journal of Physical Chemistry 2015
Molecular representation of multilayered Gram-negative bacterial membrane. The cell envelope of Gram-negative bacteria is a complex multilayered structure that, besides gatekeeping chemical transport in and out of the bacterial cell, protects the cell from hostile environments. The image shows an illustrative model of the Gram-negative bacterial membrane with lipopolysaccharide enriched outer membrane, containing O-antigen domains (pink), outer membrane proteins (in green, red, purple, yellow cartoon representation), and the phospholipid (orange and white) inner membrane. Until now, molecular level understanding of the bacterial membrane, especially over long time scales, has been challenging due to the enormous computational cost associated with molecular simulations of such complex structures. Here, we have developed and benchmarked a coarse grained force field for the lipopolysaccharide macromolecule with O-antigen domain that will enable long timescale molecular simulations of bacterial membranes with fractional computational cost.

Journal of Chemical Theory and Computation 2015
The optical properties of quantum dots are influenced by the presence of surface ligands. Here we investigate the effect of protein corona formation on the optical properties of the firefly Luciferase-CdSe quantum dot complex. To make the computational cost tractable for a large protein-quantum dot system, a multilevel approach was developed where the electronically excited quantum dot was treated quantum mechanically (QM) using electron-hole quasiparticle representation and the protein corona was represented using the molecular mechanics (MM) force field.

Langmuir 2012
Concerns of nanotoxicity caused by nanoparticle-cell interactions are becoming increasingly important as the applications of nanoparticles continue to grow. The cover illustrates the interaction of charged cone-, cube-, rod-, rice-, pyramid-, and sphere-shaped nanoparticles with a model cell membrane. High-performance molecular dynamics simulations were employed to calculate the potential of mean force associated with nanoparticle translocation as well as translocation rate constants. The results indicate that the translocation rate can vary greatly based on the particle surface charge and shape. The simulations also unravel a fascinating electrostatics-driven reorientation mechanism of faceted nanoparticles that facilitates instantaneous transport resulting in the disruption of the self-assembly of the membrane lipids

The Journal of Physical Chemistry 2010
Land masses abundant in silicate minerals surrounded by an ocean of water. Interfacial silica–water chemistry involves competing dissolution, precipitation, and polymerization reactions. These reactions are ubiquitous on Earth’s surface and occur over geological time scales. A new time-independent advanced Monte Carlo algorithm has been developed that provides insight into reaction mechanisms, equilibrium properties, and surface topography. In the present application of this method, the role of intrasurface hydrogen bonding in the dissolution process is explored.

The Journal of Physical Chemistry 2006
Potential Energy Surfaces for Photodissociation of Ammonia through a Conical Intersection Special Issue honoring Donald G. Truhlar, J. Physical Chemistry A 110, (2006).
